Getting Started¶
mzmine is an open-source, cross-platform software for mass spectrometry (MS) data processing and visualization. It supports LC-MS, LC-IMS-MS, GC-MS, MS imaging, MRM, and many more workflows - from raw data import through feature detection, alignment, annotation, statistics, and export. This page walks you through installation, initial setup, and your first steps toward data analysis.
If you prefer watching a quick mzmine introduction video, check out the Learners corner or one of the following videos:
- General introduction to mzmine with LC-MS Data & mzwizard: From Basics to Advanced Features
- GC-EI-MS, Multimodal-MS Data Processing, MALDI-MS imaging, Lipidomics
- Ion Mobility-MS Data Processing, PFAS, Contaminants
- Streamlining MS Library Generation in mzmine matching the library building workflow
Download & Installation / Updates¶
Download platform-specific installers or portable versions from GitHub Releases (see system requirements). mzmine uses semantic versioning (major.minor.patch, e.g., 4.9.0). The latest release is always at:
https://github.com/mzmine/mzmine/releases/latest
Install using the system-specific installer (e.g., .msi on Windows) by double-clicking it.
To update, install the newer version on top of the existing installation — no uninstall is required.
Portable versions are distributed as ZIP archives; extract them and run the executable directly.
Platform-specific installation guides:
No Java installation required
mzmine ships with its own specialized Java runtime. Your local Java installation has no impact on mzmine and does not need to be updated or installed.
Security warnings on first launch
Windows and macOS users may see an "untrusted source" warning the first time they run mzmine. This is expected — click Run anyway (Windows) or Open (macOS) to continue.
Running mzmine¶
mzmine provides a graphical user interface (GUI) designed for interactive data exploration, workflow optimization, and results validation. Once a workflow is optimized and no interactive review is needed, mzmine can be run headlessly as a command-line tool (CLI) for automated or server-side batch processing.
Familiarize yourself with the main window overview to understand how to navigate raw data files, processed feature lists, and visualization panels.
Sign In / Sign Up¶
On first launch, open Users → User Management to sign in to an existing account or register a free user account. See support options.
Learn more about user accounts
Configure Preferences¶
Before starting your first project, configure a few key settings:
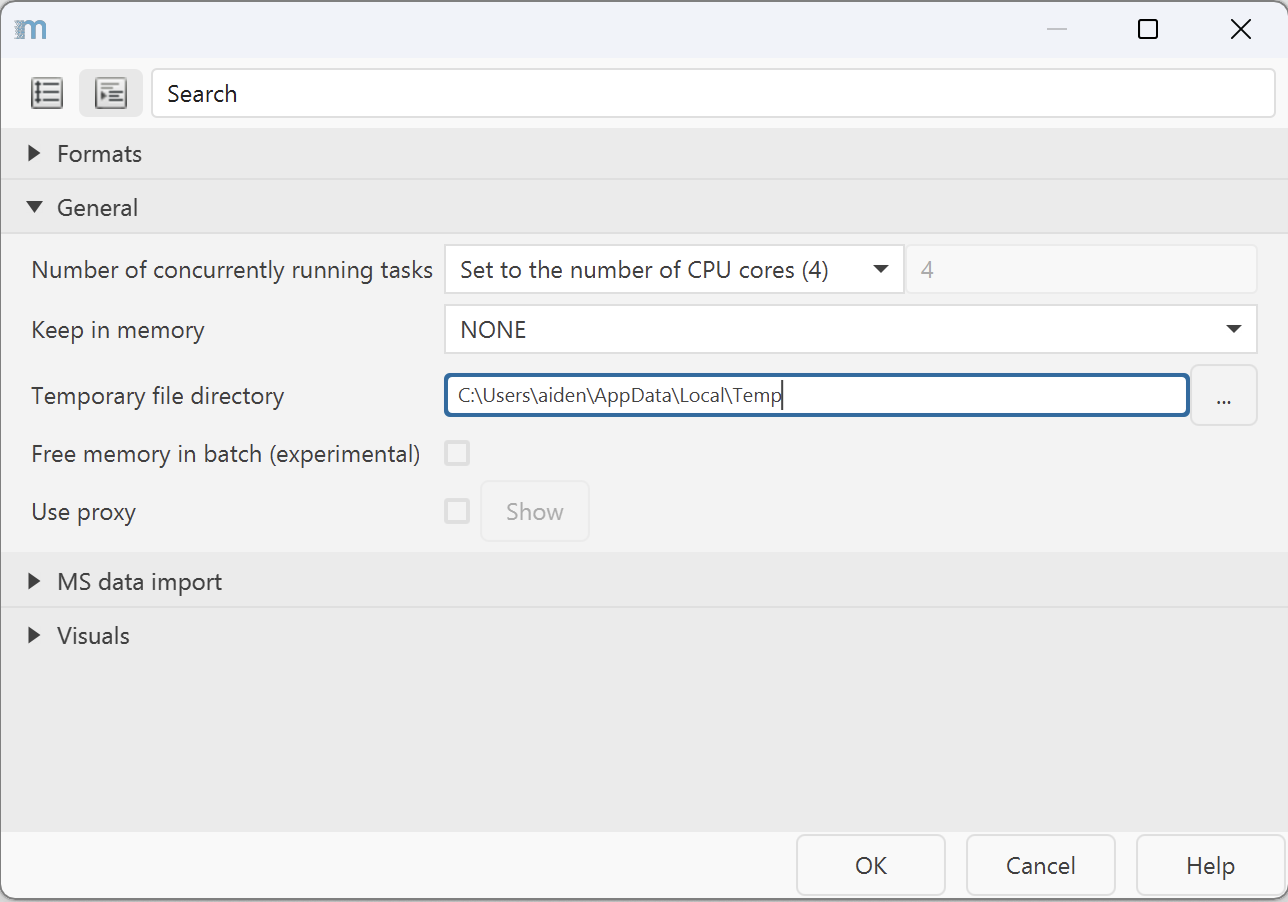
- Set a temporary file directory: go to Project → Set preferences → Temporary file
directory and restart mzmine to apply the change.
- Use a fast SSD (not your system drive) with sufficient free space for best processing performance and visualizations.
Project compatibility
mzmine 2 and mzmine 3 project files and batch files cannot be imported due to fundamental changes in data structure and parameter formats. Start a new project in mzmine 4.
Data Conversion¶
Many vendor-specific raw data formats are directly read into mzmine while other may require conversion to open formats. Supported open formats include mzML, mzXML (prefer mzML), and imzML (MS imaging).
See the data conversion guide for tools and step-by-step instructions. An option is to install msconvert and to let mzmine automatically decide to either load data natively from vendor formats or to convert in the background.
Quick Start with the Processing Wizard (mzwizard)¶
The fastest way to get started is the processing wizard (mzwizard), accessible from the main menu or the mzmine landing page. The wizard guides you through instrument-type selection, parameter configuration, and full workflow execution with minimal effort — ideal for standard LC-MS, LC-IMS-MS, GC-MS, and MS imaging datasets.
Workflows¶
Ready to process? Choose a workflow matching your instrument setup and analytical goals:
| Workflow | Description |
|---|---|
| LC-MS | Untargeted liquid chromatography–MS |
| LC-IMS-MS | Liquid chromatography–ion mobility–MS |
| GC-EI-MS | Gas chromatography–MS (EI-based) |
| MS Imaging | MALDI and other MS imaging workflows |
| MRM | Multiple reaction monitoring (targeted quantitation) |
| Library Generation | Build custom spectral libraries |
For automating and reproducing complete pipelines, see batch processing.
Learning Resources¶
New to mzmine or mass spectrometry data processing? These resources will help you get up to speed:
- Learners corner — curated video tutorials, webinars, and metabolomics courses
- mzio YouTube channel
- Workshops — recorded and upcoming training events