Features
General Features
Raw data file formats
MZmine 2 can read and process both unit mass resolution and exact mass resolution (e.g. FTMS) data in both continuous and centroided modes, including fragmentation (MSn) scans. Supported data formats are:- mzML (mzML versions 1.0 and 1.1)
- mzXML (mzXML versions 2.0 to 3.1)
- mzData (mzData versions 1.04 and 1.05)
- NetCDF (no MSn support)
- Thermo RAW
- Waters RAW
Import and Export
In order to exchange data with other applications, the following import and export file formats are supported:
Import
- CASMI challenge task
- mzTab
- XML
Export
- CSV
- MetaboAnalyst
- mzTab
- SQL database
- XML
Batch Processing
MZmine 2 has the ability to run multiple data processing methods in batch mode using any of the methods available in MZmine. The batch mode can be run as a command-line application which makes it easy to integrate MZmine into automated data analysis workflows like QC systems.Raw Data Methods
Filtering & Smoothing
There are several filtering and smoothing modules available for e.g. baseline correction, scan smoothing or cropping of the data.Peak Detection
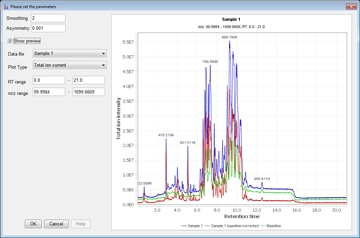
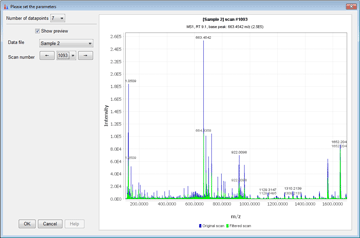
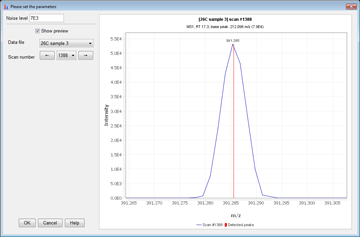
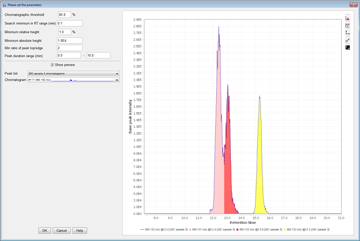
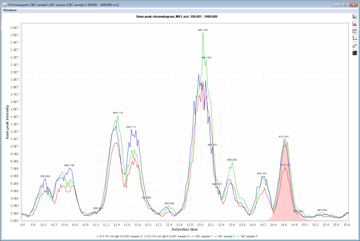
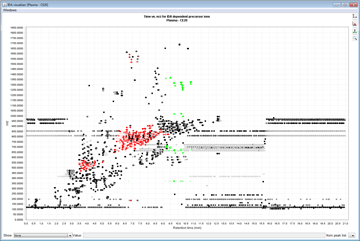
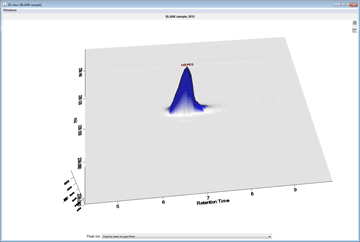
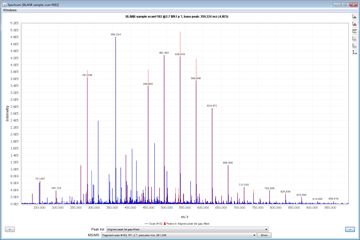
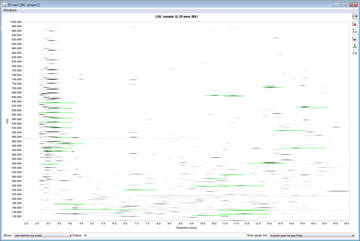
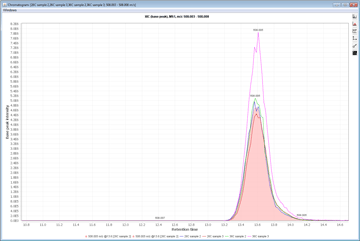
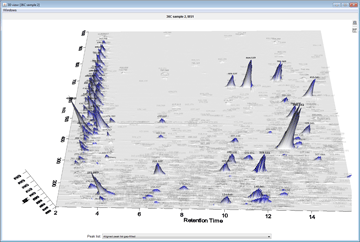
Peak detection is performed in a three-step manner: first, mass values are detected within each spectrum (several methods are available, depending on the nature of the data). In the second step, a chromatogram is constructed for each of the mass values which span over a certain time range. Finally, deconvolution algorithms are applied to each chromatogram to recognize the actual chromatographic peaks.Peak List Methods
There are several modules for processing of peaks lists, including:- Gap Filling
- Isotope Detection
- Filtering
- Alignment
- Normalization
- Identification
Identification of peaks can be performed by using any of the following:
- Custom database search
- Online database search (see below)
- Fragment or adduct search
- CAMERA
- NIST MS Search
- Various prediction tools
Currently the following online databases are supported. Support for other databases may be implemented as additional plugins.
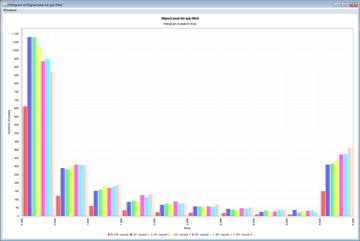
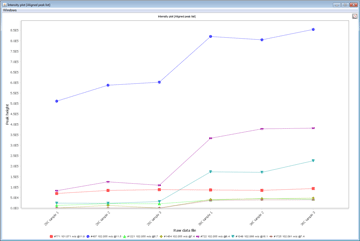
Statistical Analysis
MZmine 2 contains basic methods for statistical analysis of processed data. Processed data can also easily be exported to third-party statistical software for advanced analysis.- CV Plots
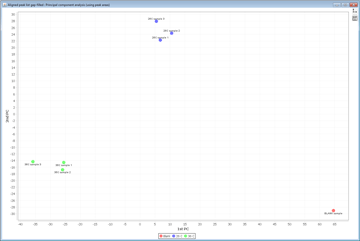
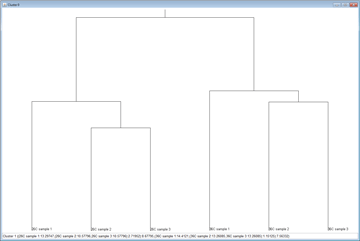
- Cluster Analysis
- Curvilinear Distant Analysis (CDA)
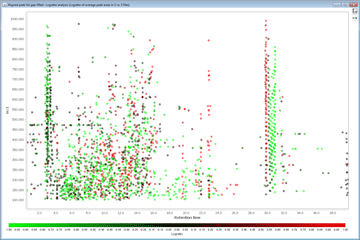
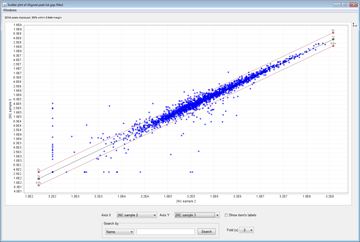
- Logratio Plots